fasdf@sadf.tu
Improved Food Safety and Public Health by Using Whole Genome Sequence Analysis
The Meeting A public meeting is being hosted by the Food Safety and Inspection Service (FSIS), the Food and Drug Administration (FDA), the Centers for Disease Control and Prevention (CDC), and the National Center for Biotechnology Information (NCBI) on Thursday and Friday, October 26 and 27, 2017. The purpose of the meeting is to protect consumers from foodborne illnesses in the USA and countries all over the world by laying the foundation for the use of whole-genome sequencing (WGS). Why Whole-Genome Sequencing (WGS) WGS analyses can determine the relationship between different bacterial isolates with higher resolution than other analytical methods, and...
fasdf@sadf.tu
Cyclospora Infections are up this Summer Sickening Nearly a 1,000 Persons in 36 States
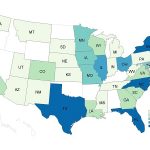
The CDC Report The CDC reported that since the beginning of 2017 to mid-September there have been 988 Laboratory confirmed cases of Cyclospora infected people in 36 states of the USA. The number of cases this year is significantly higher than in 2016. The states with the most cases are Texas 28.8%, Florida 12.0%, and New York (including NYC) 10.6%. Historical View While the number of cases is higher this year, the director of CDC’s Division of Parasitic Disease, Dr. Monica Parise, said: “The numbers from this year were probably not outside the range that we’ve seen for the last five...
fasdf@sadf.tu
FDA Commissioner Announced a Four Year Delay in Implementing Some Produce FSMA Rules
In a speech in front of National Association of State Departments of Agriculture (NASDA), Dr. Scott Gottlieb,the FDA commissioner outlined some immediate steps to facilitate the implementation of the Produce Safety Rule established by the FDA Food Safety Modernization Act (FSMA). Dr. Gottlieb claims that since being in his post, he gained a deeper appreciation for the challenges and complexity that the globalized farming community is facing. Agricultural Water Compliance Dates He announced that the FDA issued a proposed rule that, if finalized, would extend the compliance dates for the agricultural water requirements by an additional two to four years (for...
fasdf@sadf.tu
Updates: Botulism in Nacho Cheese, Salmonella in Maradol Papaya, Listeria in the Environment, and Our New Trending Page
California Department of Public Health confirms that botulism outbreak was Linked to Retail Practices In May we reported on a Botulism outbreak resulting from the consumption of nacho cheese sauce served at the Valley Oak Food and fuel gas station in Walnut Grove, California. The outbreak included 10 cases of laboratory confirmed C. botulinum toxin type A. All patients were hospitalized and one death was reported. Customers spread the nacho cheese sauce on chips from a counter-top self-service warming dispenser. According to a memo from the California Department of Public Health, the operators at the gas station were mainly responsible for the outbreak because...